Contact
Email:
mokranjac@bio.lmu.de
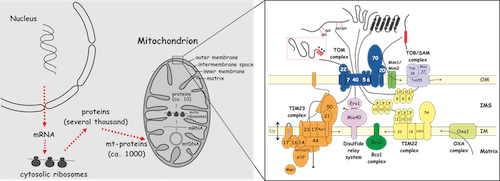 We are fascinated by mitochondria and, in particular, how they are made. Mitochondria contain their own genome and a complete apparatus for its expression, however, this genome encodes only for a handful of mitochondrial proteins. The vast majority of over thousand different mitochondrial proteins are encoded in the nuclear genome, synthesized on cytosolic ribosomes as precursor proteins with specific targeting signals and subsequently imported into the organelle. About 70% of mitochondrial proteins are synthesized with N-terminal extensions called presequences and are translocated into mitochondria with the help of the TOM and TIM23 complexes in the outer and inner membrane, respectively, along the so-called presequence pathway. We combine biochemistry with cell biology and yeast genetics and every now and then dwell into biophysics and structural biology to reveal the molecular mechanisms of protein translocation along the presequence pathway and how it is integrated into the cell physiology. The majority of our research is centered around three main questions:
We are fascinated by mitochondria and, in particular, how they are made. Mitochondria contain their own genome and a complete apparatus for its expression, however, this genome encodes only for a handful of mitochondrial proteins. The vast majority of over thousand different mitochondrial proteins are encoded in the nuclear genome, synthesized on cytosolic ribosomes as precursor proteins with specific targeting signals and subsequently imported into the organelle. About 70% of mitochondrial proteins are synthesized with N-terminal extensions called presequences and are translocated into mitochondria with the help of the TOM and TIM23 complexes in the outer and inner membrane, respectively, along the so-called presequence pathway. We combine biochemistry with cell biology and yeast genetics and every now and then dwell into biophysics and structural biology to reveal the molecular mechanisms of protein translocation along the presequence pathway and how it is integrated into the cell physiology. The majority of our research is centered around three main questions:
1. How do the TOM and TIM23 complexes cooperate during translocation of proteins?
2. How does the TIM23 complex manage to translocate some proteins across and insert the others into the inner membrane?
3. How does the TIM23 complex convert the energy of ATP hydrolysis into unidirectional transport of proteins into mitochondria?
Though most of our knowledge on the biogenesis of mitochondria comes from studies with baker´s yeast Saccharomyces cerevisiae, all the components so far identified are highly evolutionary conserved all the way to humans. Indeed, recent studies performed in mammalian systems nicely recapitulated the findings made with yeast, strengthening the use of this model system to understand the basic cellular processes in humans.
More info: https://www.cellbiology.bio.lmu.de/research_groups/mokranjac/index.html


